Arabidopsis thaliana (Thale Cress)
| Arabidopsis thaliana | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | ||||||||||||||
| Scientific classification | ||||||||||||||
| ||||||||||||||
| Binomial name | ||||||||||||||
| Arabidopsis thaliana (L.) Heynh. | ||||||||||||||
| Synonyms | ||||||||||||||
|
Arabis thaliana |
Arabidopsis thaliana is a small flowering plant widely used by plant biologists as a model organism for basic research. It has several common names including thale cress, mouse-ear cress or just Arabidopsis).[1] It is popular for research since it has a relatively short life cycle and its genome is one of the smaller ones. Due to its prominence in research it was the first plant genome to be sequenced.[2]
Habitat, morphology, and life cycle
Arabidopsis is native to Europe, Asia, and northwestern Africa. It is an annual (rarely biennial) plant usually growing to 20–25 cm tall. The leaves form a rosette at the base of the plant, with a few leaves also on the flowering stem. The basal leaves are green to slightly purplish in colour, 1.5–5 cm long and 2–10 mm broad, with an entire to coarsely serrated margin; the stem leaves are smaller, unstalked, usually with an entire margin. Leaves are covered with small unicellular hairs (called trichomes). The flowers are 3 mm in diameter, arranged on an inflorescence with a helical phyllotaxy; their structure is that of the typical Brassicacaea. The fruit is a silique 5–20 mm long, containing 20–30 seeds. Roots are simple in structure, with a single primary root that grows vertically downwards, later producing smaller lateral roots.[3][4][5][6]
Arabidopsis can complete its entire life cycle in six weeks (see the catalog subpage). The central stem that produces flowers grows after about three weeks, and the flowers naturally self-pollinate. In the laboratory Arabidopsis may be grown in petri plates or pots, under fluorescent lights or in a greenhouse.[7]
Use as a model organism
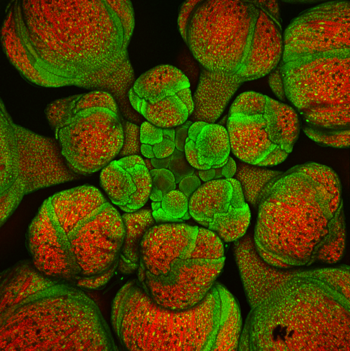
Arabidopsis is widely used as one of the model organisms for studying plant sciences, including genetics and plant development.[8][9] It plays the role for agricultural sciences that mice and fruit flies (Drosophila) play in animal biology. Although Arabidopsis thaliana has little direct significance for agriculture, it has several traits that make it a useful model for understanding the genetic, cellular, and molecular biology of flowering plants.
The small size of its genome make Arabidopsis thaliana useful for genetic mapping and sequencing — with about 157 million base pairs[10] and five chromosomes, Arabidopsis has one of the smallest genomes among plants. It was the first plant genome to be sequenced, completed in 2000 by the Arabidopsis Genome Initiative.[2] Much work has been done to assign functions to its 27,000 genes and the 35,000 proteins they encode.[11]
The plant's small size and rapid life cycle are also advantageous for research. Having specialized as a spring ephemeral, it has been used to found several laboratory strains that take about six weeks from germination to mature seed. The small size of the plant is convenient for cultivation in a small space and it produces many seeds. Further, the selfing nature of this plant assists genetic experiments. Also, as an individual plant can produce several thousand seeds, each of the above criteria leads to Arabidopsis thaliana being valued as a genetic model organism.
Finally, plant transformation in Arabidopsis is routine, using Agrobacterium tumefaciens to transfer DNA to the plant genome. The current protocol, termed "floral-dip", involves simply dipping a flower into a solution containing Agrobacterium, the DNA of interest, and a detergent.[12] This method avoids the need for tissue culture or plant regeneration.
History of Arabidopsis research
The first mutant in Arabidopsis was documented in 1873 by Alexander Braun, describing a double flower phenotype (the mutated gene was likely Agamous, cloned and characterized in 1990).[13] However, it was not until 1943 that Friedrich Laibach (who had published the chromosome number in 1907) proposed Arabidopsis as a model organism.[14] His student Erna Reinholz published her thesis on Arabidopsis in 1945, describing the first collection of Arabidopsis mutants that they generated using x-ray mutagenesis. Laibach continued his important contributions to Arabidopsis research by collecting a large number of ecotypes. With the help of Albert Kranz, these were organised into the current ecotype collection of 750 natural accessions of Arabidopsis thaliana from around the world.
In the 1950s and 1960s John Langridge and George Rédei played an important role in establishing arabidopsis as a useful organism for biological laboratory experiments. Rédei wrote several scholarly reviews instrumental in introducing the model to the scientific community. The start of the arabidopsis research community dates to a newsletter called Arabidopsis Information Service (AIS), established in 1964. The first International Arabidopsis Conference was held in 1965, in Göttingen, Germany.
In the 1980s Arabidopsis started to become widely used in plant research laboratories around the world. It was one of several model organisms, that included maize, petunia and tobacco, used for basic research.[14] The latter two were attractive since they were easily transformable with the then current technologies, while maize was a well established genetic model for plant biology. The breakthrough year for Arabidopsis as the preferred model plant came in 1986 when T-DNA mediated transformation was first published and this coincided with the first gene to be cloned and published.[15][16]
References
- ↑ Germplasm Resources Information Network: Arabidopsis thaliana
- ↑ 2.0 2.1 The Arabidopsis Genome Initiative (2000). "Analysis of the genome sequence of the flowering plant Arabidopsis thaliana". Nature 408: 796–815. DOI:10.1038/35048692. PMID 11130711. Research Blogging.
- ↑ Flora of NW Europe: Arabidopsis thaliana
- ↑ Blamey, M. & Grey-Wilson, C. (1989). Flora of Britain and Northern Europe. ISBN 0-340-40170-2
- ↑ Flora of Pakistan: Arabidopsis thaliana
- ↑ Flora of China: Arabidopsis thaliana
- ↑ D.W. Meinke, J.M. Cherry, C. Dean, S.D. Rounsley, M. Koornneef (1998). "Arabidopsis thaliana: A Model Plant for Genome Analysis". Science 282 (5389): 662–682. DOI:10.1126/science.282.5389.662. Research Blogging.
- ↑ Rensink WA, Buell CR (2004). "Arabidopsis to rice. Applying knowledge from a weed to enhance our understanding of a crop species". Plant Physiol. 135 (2): 622–9. DOI:10.1104/pp.104.040170. PMID 15208410. Research Blogging.
- ↑ Coelho SM, Peters AF, Charrier B, et al (2007). "Complex life cycles of multicellular eukaryotes: new approaches based on the use of model organisms". Gene 406 (1-2): 152–70. DOI:10.1016/j.gene.2007.07.025. PMID 17870254. Research Blogging.
- ↑ Bennett, M. D., Leitch, I. J., Price, H. J., & Johnston, J. S. (2003). "Comparisons with Caenorhabditis (100 Mb) and Drosophila (175 Mb) Using Flow Cytometry Show Genome Size in Arabidopsis to be 157 Mb and thus 25% Larger than the Arabidopsis Genome Initiative Estimate of 125 Mb". Annals of Botany 91: 547–557. DOI:10.1093/aob/mcg057. PMID 12646499. Research Blogging.
- ↑ Integr8 - A.thaliana Genome Statistics:.
- ↑ Zhang X, Henriques R, Lin SS, Niu QW, Chua NH (2006). "Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method". Nat Protoc 1 (2): 641–6. DOI:10.1038/nprot.2006.97. PMID 17406292. Research Blogging.
- ↑ M.F. Yanofsky, H. Ma, J.L. Bowman, G.N. Drews, K.A. Feldmann & E.M. Meyerowitz (1990). "The protein encoded by the Arabidopsis homeotic gene agamous resembles transcription factors". Nature 346: 35–39. DOI:10.1038/346035a0. PMID 1973265. Research Blogging.
- ↑ 14.0 14.1 E.M. Meyerowitz (2001). "Prehistory and History of Arabidopsis Research". Plant Physiology 125: 15–19. DOI:10.1038/346035a0. PMID 11154286. Research Blogging.
- ↑ Lloyd AM, Barnason AR, Rogers SG, Byrne MC, Fraley RT, Horsch RB (1986). "Transformation of Arabidopsis thaliana with Agrobacterium tumefaciens". Science 234: 464–466. DOI:10.1126/science.234.4775.464. PMID 17792019. Research Blogging.
- ↑ Chang C, Meyerowitz EM (1986). "Molecular cloning and DNA sequence of the Arabidopsis thaliana alcohol dehydrogenase gene". Proc Natl Acad Sci USA 83: 1408–1412. DOI:10.1073/pnas.83.5.1408. PMID 2937058. Research Blogging.