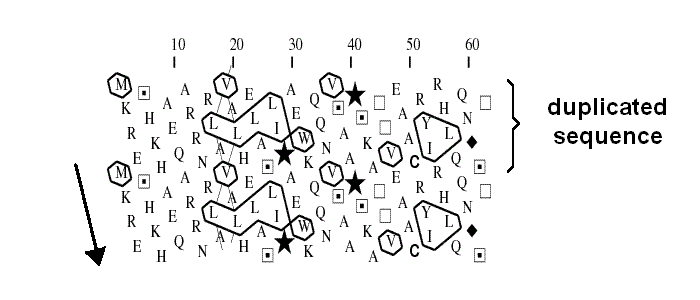
Hydrophobic cluster analysis
Amino acid sequences with very similar distribution patterns of the hydrophobic set of residues V, I, L, F, Y, W, M (detected in a two-dimensional helical representation of the protein sequence) are most often structural homologs, even when the overall sequence identity is as low as 7 %. This representation is obtained by writing the protein sequence on a classical alpha helix (3.6 amino acids per turn) smoothed on a cylinder. After five turns, residues i and i+ 18 have similar positions parallel to the axis of the cylinder. To make this 3D representation easier to handle, the cylinder is then cut parallel to its axis and unrolled. As some adjacent amino acids are widely separated by the unfolding of the cylinder, the representation is duplicated, making the sequence easier to follow and giving a better impression of the environment of each aminoacid.
Clusters of these hydrophobic amino acids are good markers of regular secondary structures and have been extensively used in the detection of similar folds or similar motifs between sequences showing very limited sequence relatedness.
References
- Gaboriaud C, Bissery V, Benchetrit T and Mornon JP (1987) Hydrophobic cluster analysis: an efficient new way to compare and analyse amino acid sequences. FEBS Lett. 224, 149-155
- Hennetin J, Tuan KL, Canard L, Colloc’h N, Mornon JP, Callebaut I (2003) Non-intertwined binary patterns of hydrophobic/nonhydrophobic amino acids are considerably better markers of regular secondary structures than nonconstrained patterns. Proteins 51, 236-244
- Callebaut I, Labess G, Durand P, Poupon A, Canard L, Chomilier J, Henrissat B, Mornon JP (1997) Deciphering protein sequence information through hydrophobic cluster analysis (HCA): current status and perspectives. Cell Mol Life Sci 53, 621-645